OVERVIEW
Organisms that reproduce sexually utilize a specialized cell division program called meiosis to reduce their chromosome number by half to generate haploid gametes (sperm and eggs). Proper execution of this process is crucial, since errors in meiotic chromosome segregation result in aneuploidy, a leading cause of miscarriages and birth defects in humans. Female meiosis in particular is highly error prone and this vulnerability has a profound impact on human health; it is estimated that as many as 10-25% of human embryos are chromosomally abnormal, and the vast majority of these defects arise from problems with the female reproductive cells (oocytes). However, the molecular mechanisms driving these important divisions are not well-understood.
Research in the Wignall lab is aimed at exploring how oocytes divide. We are tackling this problem through studies in the model organism C. elegans, where we use a variety of cell biological, genetic, and biochemical approaches, including state-of-the-art imaging, CRISPR-mediated mutagenesis, methods for rapid protein depletion, and in vitro reconstitution assays.
PROJECTS
Our research is focused in two major areas:
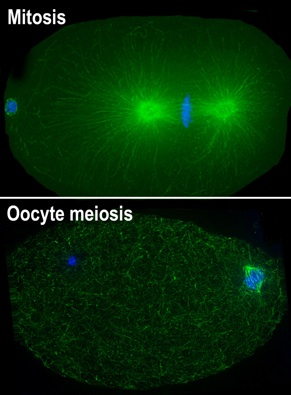
(1) We study a dynamic macromolecular machine, the spindle, which physically separates the chromosomes during cell division. We are interested in understanding how this intricate structure forms and is stabilized, so that it can properly function to partition the genetic material. Notably, oocytes lack centrosomes, which nucleate microtubules and act as structural cues to define and organize the spindle poles in other cell types. Consequently, oocyte spindles are morphologically distinct and it is therefore important to study oocytes directly to understand how these acentrosomal spindles form and mediate accurate chromosome partitioning.
(2) To be partitioned correctly, chromosomes must attach to the spindle. In most cells, these attachments are formed when microtubules associate with chromosomal structures called kinetochores. However, we previously found that chromosomes in C. elegans oocytes do not form canonical kinetochore attachments during the period where they are moving to the center of the spindle, and instead, microtubules run laterally alongside chromosomes. We study how chromosome-associated proteins mediate this unique form of attachment and drive chromosomal movements during cell division.
Altogether, our goal is to advance our understanding of how genomic integrity is maintained during cell division.
