Metal Regulation
Microbes use a diverse set of metal-specific sensors known as metalloregulatory proteins to respond to changes in environmental metal concentrations. These transcription factors control expression of many diverse factors including membrane bound metal ion transporters, or metallochaperones, that optimize cellular physiology in response to dynamic shifts in metal availability.
Zinc Circuitry
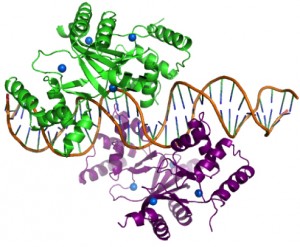
Figure 1: Structure of two Zur dimers bound to DNA.
One Zur dimer (green) binds to the Zr DNA site (orange). A second Zur dimer (purple) binds to the adjacent face of the Zur DNA binding site. Each Zur dimer binds four zinc ions (blue). (Gilston et al., 2014).
Zinc uptake regulator, Zur, is a Fur (ferric uptake regulator) family transcription repressor of the Zur regulon in E. coli. Zur is extremely sensitive to changes in zinc cellular zinc concentrations suggesting that bacteria are intolerant to of free zinc in the cytosol under normal growth conditions. To understand how microorganisms coordinate this dynamic response to changes in zinc availability at the molecular level, the O’Halloran group solved the crystal structure of Zur bound to DNA (Figure 1). The group’s results provide strong support for the idea that the thermodynamics of an ensemble of protein–DNA interactions controls the zinc-sensing gene regulation network through co-operative binding of Zur dimers to adjacent faces of DNA. This cooperative of binding of Zur dimers leads to the differential regulation patters in this regulon. Through bioinformatics and thermodynamic, biochemical and physiological assays, the group also identified the promoter of the periplasmic lysozyme inhibitor gene (pliG) as a new novel target of Zur.
Selected Publications:
• Gilston BA, Wang S, Marcus MD, Canalizo-Hernandez MA, Swindell EP, Xue Y, Mondragon A, O’Halloran TV. Structural and mechanistic basis of zinc regulation across the E. coli Zur regulon. PLoS Biology. 2014;12(11):e1001987.
• Outten, C.E., Tobin, D.A., Penner-Hahn, J.E., O’Halloran, T.V. “Characterization of the Metal Receptor Sites in E. coli Zur, an Ultrasensitive Zn(II) Metalloregulatory Protein.” Biochemistry, 2001, 40, 35, 10417-10423.
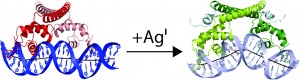
Figure 2: CueR alters the topology of DNA.
In the absence of metal, CueR dimer (red) binds to a 33 bp fragment of the copA promoter to repress transcription. Addition of metal switches CueR to the activator state (green). In the activator state, CueR changes the topology of copA to promote binding of RNA polymerase. (Philips et al., 2015).
Copper Circuitry
CueR, a member of the MerR protein family, responds to zeptomolar concentrations of free copper and is the most sensitive of the copper sensors in enterobacteria. Members of the MerR metalloregulatory protein family exert tight control over transcription and act at a single site within the prokaryotic core promoter both as repressors and as ligand-inducible activators of transcription. The O’Halloran group determined the structure of CueR/DNA with and without bound metal to resolve the paradox of how CueR can both repress and activate transcription. Crystal structures of the CueR-DNA complexes reveal how the repressor and activator torque and bend DNA in this genetic switching event to alter the distance between the -35 and -10 promoter elements to prevent or promote binding of RNA polymerase (Figure 2). It is this new overall topology that activates transcription, rather than re-phasing of the -10 and -35 promoter elements.
Selected Publications:
• Philips SJ, Canalizo-Hernandez M, Yildirim I, Schatz GC, Mondragón A, O’Halloran TV. “Allosteric transcriptional regulation via changes in the overall topology of the core promoter”. Science. 2015 Aug 21;349(6250):877-81.
• Outten, F.W., Outten, C., Hale, J., and O’Halloran, T. “Transcriptional Activation of an E. coli Copper Efflux Regulon by the Chromosomal MerR Homologue, CueR.” J. Biological Chemistry, 2000, 275 (40) 31024-31029.
• Ansari AZ, Bradner JE, O’Halloran TV. DNA-bend modulation in a repressor-to-activator switching mechanism. Nature, 1995, 374(6520):371-5.
• Ansari, A.Z., Chael, M.L., O’Halloran, T.V. “Allosteric Underwinding of DNA is a Critical Step in Positive Control of Transcription by MerR.” Nature, 1992, 355, 87-89.