Northwestern University researchers have successfully coaxed a deadly pathogen to destroy itself from the inside out.
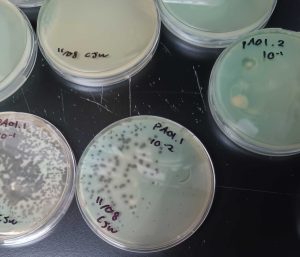
The dark spots in the dishes mark areas where phage burst out of the bacteria, killing them. Photo credit: Cole Wilson
In the new study, researchers modified DNA from a bacteriophage or “phage,” a type of virus that infects and replicates inside of bacteria. Then, the research team put the DNA inside Pseudomonas aeruginosa (P. aeruginosa), a deadly bacterium that is also highly resistant to antibiotics. Once inside the bacterium, the DNA bypassed the pathogen’s defense mechanisms to assemble into virions, which sliced through the bacterium’s cell to kill it.
Building on a growing interest in “phage therapies,” the experimental work represents a critical step toward engineering designer viruses as new therapeutics to kill antibiotic-resistant bacteria. It also reveals vital information about the innerworkings of phages, a little-studied area of biology.
The study was published Jan. 31 in the journal Microbiology Spectrum.
“Antimicrobial resistance is sometimes referred to as the ‘silent pandemic,’” said Northwestern’s Erica Hartmann, who led the work. “The numbers of infections and deaths from infections are increasing worldwide. It’s a massive problem. Phage therapy has emerged as an untapped alternative to our reliance on using antimicrobials. But, in many ways, phages are microbiology’s ‘final frontier.’ We don’t know much about them. The more we can learn about how phage work, the more likely we can engineer more effective therapeutics. Our project is cutting-edge in that we are learning about phage biology in real time as we engineer them.”
An indoor microbiologist, Hartmann is an associate professor of civil and environmental engineering at Northwestern’s McCormick School of Engineering and a member of the Center for Synthetic Biology.
Desperate need for antibiotic alternatives
Associated with an increase in antimicrobial use, the rise of antibacterial resistance is an urgent and growing threat to the global population. According to the Centers for Disease Control and Prevention (CDC), nearly 3 million antimicrobial-resistant infections occur each year in the United States alone, with more than 35,000 people dying as a result.
The growing crisis has motivated researchers to look for alternatives to antibiotics, which are continually losing effectiveness. In recent years, researchers have started to explore phage therapies. But even though billions of phages exist, scientists know very little about them.
“For every bacterium that exists, there are dozens of phages,” Hartmann said. “So, there is an astronomically large number of phages on Earth, but we only understand a handful of them. We haven’t necessarily had the motivation to really study them. Now, the motivation is there, and we are increasing the number of tools we have to dedicate to their study.”
Treatment without side effects
To explore potential phage therapies, researchers either pinpoint or modify an existing virus to selectively target a bacterial infection without disrupting the rest of the body. Ideally, scientists one day could tailor a phage therapeutic to infect a specific bacterium and design “a la carte” therapeutics with precise traits and characteristics to treat individual infections.
“What’s powerful about phage is it can be very specific in the way that antibiotics are not,” Hartmann said. “If you take an antibiotic for a sinus infection, for example, it disrupts your entire gastrointestinal tract. A phage therapy can be designed to affect only the infection.”
While other researchers have investigated phages therapies, almost all of those studied have focused on using phages to infect Escherichia coli. Hartmann, however, decided to focus on P. aeruginosa, one of the five most deadly human pathogens. Particularly dangerous for people with compromised immune systems, P. aeruginosa is a leading cause of hospital infections, often infecting patients with burn or surgery wounds as well as lungs in people with cystic fibrosis.
“It is one of the highest priority, multi-drug resistant pathogens that many people are really concerned about,” Hartmann said. “It is extremely drug resistant, so there is an urgent need to develop alternative therapeutics for it.”
Mimicking infection, bypassing defenses
In the study, Hartmann and her team started with P. aeruginosa bacteria and purified DNA from several phages. Then, they used electroporation — a technique that delivers short, high-voltage pulses of electricity — to poke temporary holes in the bacteria’s outer cell. Through these holes, phage DNA entered the bacteria to mimic the process of infection.
In some cases, the bacteria recognized the DNA as a foreign object and shredded the DNA to protect itself. But after using synthetic biology to optimize the process, Hartmann’s team was able to knock out the bacteria’s antiviral self-defense mechanisms. In these cases, the DNA successfully carried information into the cell, resulting in virions that killed the bacteria.
“Where we were successful, you can see dark spots on the bacteria,” Hartmann said. “This is where the viruses burst out of the cells and killed all the bacteria.”
After this success, Hartmann’s team introduced DNA from two more phages that are naturally unable to infect their strain of P. aeruginosa. Yet again, the process worked.
Phage manufacturing in a cell
Not only did the phage kill the bacteria, the bacteria also ejected billions more phages. These phages can then be used to kill other bacteria, like those causing an infection.
Next, Hartmann plans to continue modifying phage DNA to optimize potential therapies. For now, her team is studying the phages expelled from the P. aeruginosa.
“This is an important piece in making phage therapies,” she said. “We can study our phage in order to decide which ones to develop and eventually mass produce them as a therapeutic.”
The study, “A synthetic biology approach to assemble and reboot clinically relevant Pseudomonas aeruginosa tailed phages,” was supported by the Walder Foundation, the National Science Foundation and the National Institutes of Health.
This story originally appeared on Northwestern Now.